
Breaking News
 Food Crisis Is Coming | MAHA's John Klar
Food Crisis Is Coming | MAHA's John Klar
 He Forked systemd to Stop Age Verification... Then the Original Dev Showed Up!
He Forked systemd to Stop Age Verification... Then the Original Dev Showed Up!
 MASSIVE RAINFALL After Iran Destroys U.S. Weather Modification Radar!
MASSIVE RAINFALL After Iran Destroys U.S. Weather Modification Radar!
 FORD'S NIGHTMARE TRUCK IS HERE - They just patented a truck that you won't own,...
FORD'S NIGHTMARE TRUCK IS HERE - They just patented a truck that you won't own,...
Top Tech News
 Researcher wins 1 bitcoin bounty for 'largest quantum attack' on underlying tech
Researcher wins 1 bitcoin bounty for 'largest quantum attack' on underlying tech
 Interceptor-Drone Arms-Race Emerges
Interceptor-Drone Arms-Race Emerges
 A startup called Inversion has introduced Arc, a space-based vehicle...
A startup called Inversion has introduced Arc, a space-based vehicle...
 Mining companies are using cosmic rays to find critical minerals
Mining companies are using cosmic rays to find critical minerals
 They regrew a severed nerve - by shortening a bone.
They regrew a severed nerve - by shortening a bone.
 New Robot Ants Work Like Real Insects To Build And Dismantle On Their Own
New Robot Ants Work Like Real Insects To Build And Dismantle On Their Own
 Russian scientists 'are developing the world's first drug to delay ageing' months after
Russian scientists 'are developing the world's first drug to delay ageing' months after
 Sam Altman's World ID Expands Biometric Identity Checks
Sam Altman's World ID Expands Biometric Identity Checks
 China Tests Directed Energy Beam That Recharges Drones Mid-Flight
China Tests Directed Energy Beam That Recharges Drones Mid-Flight
 Jurassic Park might arrive sooner than expected, just with Dinobots.
Jurassic Park might arrive sooner than expected, just with Dinobots.
Harvard scientists uncover an exploitable Achilles' heel common to most bacteria
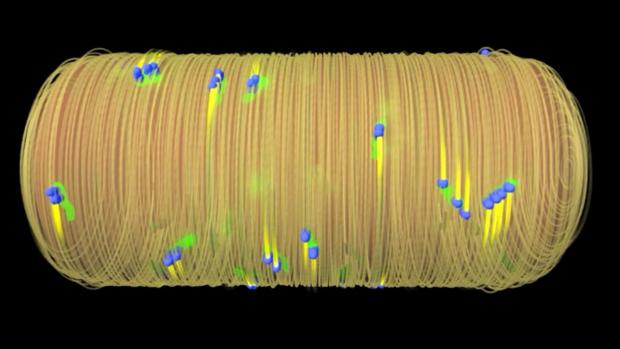
Finding ways to disarm these defenses is a key component of antibiotics, and now researchers at Harvard Medical School have identified a structural weakness that seems to be built into a range of bacterial species, potentially paving the way for a new class of widely-effective antibacterial drugs.
The new study builds on previous research into a protein named RodA. While the protein itself has long been known, in 2016 the Harvard team was the first to discover that it builds the protective cell walls of bacteria out of sugar molecules and amino acids. Since RodA belongs to the SEDS family of proteins, which is common to almost all bacteria, the team realized it was the perfect target for a far-reaching antibiotic. And on closer examination of RodA, the researchers spotted a vulnerable looking cavity on the outer surface of the protein.
"What makes us excited is that this protein has a fairly discrete pocket that looks like it could be easily and effectively targeted with a drug that binds to it and interferes with the protein's ability to do its job," says David Rudner, co-senior author of the study.
To test whether this cavity was the Achilles' heel they were looking for, the scientists altered the structure of the protein in two species of bacteria, E. coli and Bacillus subtilis. These two were chosen because they're well understood and represent the two broad classes of disease-causing bacteria, gram-positive and gram-negative.



